Visualizar Tese - Instituto de Biociências - Unesp
Visualizar Tese - Instituto de Biociências - Unesp Visualizar Tese - Instituto de Biociências - Unesp
[53] Nikitenko LL, Cross T, Campo L et al. (2006) Expression of terminally glycosylatedcalcitonin receptor-like receptor in uterine leiomyoma: endothelial phenotype and associationwith microvascular density. Clin Cancer Res 12: 5648-5658.[54] Lin N, Chen S, Pan W et al. (2011) NP603, a novel and potent inhibitor of FGFR1tyrosine kinase, inhibits hepatic stellate cell proliferation and ameliorates hepatic fibrosis inrats. Am J Physiol Cell Physiol, in press.19
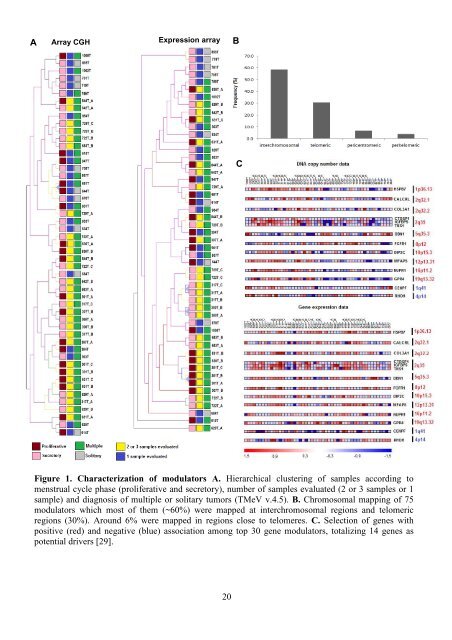
AArray CGHExpression arrayBtumoral phenotype.CFigure 1. Characterization of modulators A. Hierarchical clustering of samples according tomenstrual cycle phase (proliferative and secretory), number of samples evaluated (2 or 3 samples or 1sample) and diagnosis of multiple or solitary tumors (TMeV v.4.5). B. Chromosomal mapping of 75modulators which most of them (~60%) were mapped at interchromosomal regions and telomericregions (30%). Around 6% were mapped in regions close to telomeres. C. Selection of genes withpositive (red) and negative (blue) association among top 30 gene modulators, totalizing 14 genes aspotential drivers [29].20
- Page 241 and 242: Cirilo, PDRREFERÊNCIAS218Kazmiercz
- Page 243 and 244: Cirilo, PDRREFERÊNCIAS220Lloret M,
- Page 245 and 246: Cirilo, PDRREFERÊNCIAS222Mitra R,
- Page 247 and 248: Cirilo, PDRREFERÊNCIAS224Ozisik YY
- Page 249 and 250: Cirilo, PDRREFERÊNCIAS226Redon R,
- Page 251 and 252: Cirilo, PDRREFERÊNCIAS228Solyom S,
- Page 253 and 254: Cirilo, PDRREFERÊNCIAS230Ueki K, R
- Page 255 and 256: Cirilo, PDRREFERÊNCIAS232and poly(
- Page 257 and 258: Skubitz KM e Skubitz APJ Lab Clin M
- Page 259 and 260: Dimitrova IK et al. Fertil Steril,9
- Page 261 and 262: 317T_B317T_C13 614T 44 LU prolifera
- Page 263 and 264: 46 867T 46 LUM outros B 14 33,5 17
- Page 265 and 266: 1015T 6q13-q14.1 75,806,978-76,314,
- Page 267 and 268: MRPS31, SLC25A15, SUGT1L1 320d-1630
- Page 269 and 270: 8q22.3-q23.1 105,831,966 107,742,33
- Page 271 and 272: Anexo 5. Genes identificados na an
- Page 273 and 274: + + CCDC8 0 51 37 14+ - CALM3 51 0
- Page 275 and 276: ABSTRACTBackground: Uterine leiomyo
- Page 277 and 278: Thus, these findings prompt us to i
- Page 279 and 280: Technologies, Santa Clara, CA). Sta
- Page 281 and 282: mapped at 1p36.13, 1q41, 2q32.1, 2q
- Page 283 and 284: genetic disorder showed the higher
- Page 285 and 286: The modulators were mainly associat
- Page 287 and 288: egulation of cell cycle. Chegini an
- Page 289 and 290: REFERENCES[1] Baird DD, Dunson DB (
- Page 291: [37] Luo X, Ding L, Xu J et al. (20
- Page 295 and 296: ABDiseases and Disorders P value Mo
- Page 298 and 299: Tabel 2. Genes identified on module
- Page 300 and 301: Table S1. Clinical parameters and h
- Page 302 and 303: 32 954T 45 M secretory W 12 33,7 18
- Page 304 and 305: 16 28184420-30915100 2730680 p11.2
- Page 306 and 307: MYPN hsa-miR-214 -NFKBIL2 - -PKP3 -
- Page 308: Table S5. Ingenuity Pathways Analys
AArray CGHExpression arrayBtumoral phenotype.CFigure 1. Characterization of modulators A. Hierarchical clustering of samples according tomenstrual cycle phase (proliferative and secretory), number of samples evaluated (2 or 3 samples or 1sample) and diagnosis of multiple or solitary tumors (TMeV v.4.5). B. Chromosomal mapping of 75modulators which most of them (~60%) were mapped at interchromosomal regions and telomericregions (30%). Around 6% were mapped in regions close to telomeres. C. Selection of genes withpositive (red) and negative (blue) association among top 30 gene modulators, totalizing 14 genes aspotential drivers [29].20