note - FIZ Karlsruhe
note - FIZ Karlsruhe
note - FIZ Karlsruhe
Create successful ePaper yourself
Turn your PDF publications into a flip-book with our unique Google optimized e-Paper software.
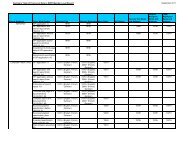
Display selected answers in a bibliographic format<br />
=> D BIB AB SCORE ALIGN<br />
Introduction to similarity searching<br />
L3 ANSWER 1 OF 8554 DGENE COPYRIGHT 2010 THOMSON REUTERS on STN<br />
AN AEF76660 DNA DGENE Full-text<br />
TI Isolated nucleic acid, coded for a fusion protein, is formed by two<br />
functionally bonded nucleic acid sequences where one is derived from a<br />
mycovirus.<br />
IN Schmitt M J; Powilleit F<br />
PA (UYSA-N) UNIV SAARLANDES.<br />
PI DE 102004032888 A1 20060202 50<br />
AI DE 2004-102004032888 20040707<br />
PRAI DE 2004-102004032888 20040707<br />
PSL Claim 13; SEQ ID NO 1<br />
DT Patent<br />
LA German<br />
OS 2006-175023 [19]<br />
CR P-PSDB: AEF76661<br />
DESC Human herpesvirus 5 pp65 encoding DNA SEQ ID NO 1.<br />
AB This invention describes a novel recombinant nucleic acid, encoding a<br />
fusion protein which can form virus-type particles and has at least one<br />
nucleic acid sequence derived from a mycovirus and a nucleic acid<br />
sequence which does not come from a mycovirus. The nucleic acid is<br />
inserted into a host cell and cultivated under conditions to form or<br />
promote recombinant virus-type particles. The products of the invention<br />
are useful for the treatment of infectious diseases, for the<br />
heterologeous expression of proteins in yeast cells, for the study of<br />
protein/protein interactions and the for the production of transfection<br />
agents. The method gives a nucleic acid which gives an expression of<br />
foreign proteins e.g. for the development of vaccines, the production of<br />
proteins and especially enzymes. This sequence encodes a protein used in<br />
the method of the invention. NOTE: The sequence data for this patent was<br />
not represented in the specification but was obtained directly from the<br />
DE Patent Office.<br />
SCORE 251 96% of query self score 260<br />
ALIGN Smith-Waterman score: 251<br />
52 na overlap starting at 2631<br />
ggguuuaggagugguaggucuuacgaugccagcuguaaugccuaccggataa<br />
:::...:::::.::.:::.:..::::.::::::.:.::.:::.:::::: ::<br />
Gggtttaggagtggtaggtcttacgatgccagctgtaatgcctaccggagaa<br />
<strong>note</strong><br />
Helpful<br />
HINT<br />
One dot represents the “family” similarity of Uracil (U) to Thymine (T).<br />
GETSIM is slower than BLAST, so using GETSIM in BATCH mode is<br />
recommended. In addition, BATCH query limits are higher. See page 79.<br />
GENESEQ on STN (DGENE) Workshop Manual | Page 39