note - FIZ Karlsruhe
note - FIZ Karlsruhe
note - FIZ Karlsruhe
You also want an ePaper? Increase the reach of your titles
YUMPU automatically turns print PDFs into web optimized ePapers that Google loves.
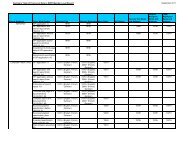
VIII: BLAST advanced options<br />
Parameter Code Defaults / Valid values<br />
Expectation value -e<br />
Filter -f<br />
Word Size -w<br />
Strand -s<br />
Matrix -m<br />
Gap Penalty -g<br />
Gap Extension -x<br />
Penalty for<br />
nucleotide mismatch<br />
Reward for<br />
nucleotide match<br />
Helpful<br />
HINT<br />
Default 10; Integers and floating point<br />
numbers<br />
Default t (true, on); additionally f (false,<br />
off), and c (coiled coiled) for nucleotides<br />
only<br />
Default 11 for nucleotides (7-23)<br />
Default 3 for peptides (2)<br />
Default 3 resp. BOTH; 1 resp. SIN and 2<br />
resp. COM<br />
Default BLOSUM62; additionally<br />
BLOSUM45, BLOSUM80, PAM30 and<br />
PAM70<br />
Default 11 for peptides<br />
Default 5 for nucleotides<br />
(positive integers)<br />
Note that for a given matrix only certain<br />
gap costs are allowed<br />
Default 1 for peptides<br />
Default 2 for nucleotides<br />
(positive integers)<br />
Note that for a given matrix only certain<br />
gap extension costs are allowed<br />
-q Default –3 (negative integers)<br />
-r Default 1 (1-8)<br />
See also Exploring BLAST advanced options on page 49.<br />
Appendices<br />
GENESEQ on STN (DGENE) Workshop Manual | Page 137