View - ResearchGate
View - ResearchGate View - ResearchGate
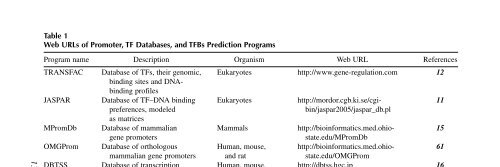
Table 1Web URLs of Promoter, TF Databases, and TFBs Prediction ProgramsProgram name Description Organism Web URL ReferencesTRANSFAC Database of TFs, their genomic, Eukaryotes http://www.gene-regulation.com 12binding sites and DNAbindingprofilesJASPAR Database of TF–DNA binding Eukaryotes http://mordor.cgb.ki.se/cgi- 11preferences, modeled bin/jaspar2005/jaspar_db.plas matricesMPromDb Database of mammalian Mammals http://bioinformatics.med.ohio- 15gene promoters state.edu/MPromDbOMGProm Database of orthologous Human, mouse, http://bioinformatics.med.ohio- 61mammalian gene promoters and rat state.edu/OMGPromDBTSS Database of transcription Human, mouse, http://dbtss.hgc.jp 16start sites zebrafish, malaria,and schyzonEPD Database of eukaryotic Eukaryotes http://www.epd.isb-sib.ch 67gene promotersUCSC Genome browser at UCSC Genome sequences http://genome.ucsc.edu 68and annotationsincluding human–mouse–ratconserved blocksMatch Program to search for TFBSs Eukaryotes http://www.gene-regulation.com/ 19using TRANSFAC PWMs pub/programs.html#matchTESS Program to predict TFBSs Eukaryotes http://www.cbil.upenn. –edu/cgi-bin/tess/tessMatInspector Program to search for TFBSs Eukaryotes http://www.genomatix.de/products/ 50using TRANSFAC PWMs MatInspector/MatInspector1.html132
- Page 236: 106 Crabtree et al.17. Some cluster
- Page 240: 108 Crabtree et al.19. Chado—The
- Page 244: 110 Dateproducts prevents the under
- Page 248: 112 DateDetails of these tasks are
- Page 252: 114 DateThis step creates additiona
- Page 256: 116 Date>hsapiens|gi|20093443 >hsap
- Page 260: 118 DateBLAST score from the match
- Page 264: Table 1A Sample of Results From Pro
- Page 268: 122 DateFig. 1. A network of functi
- Page 272: 124 Datedescribed by Verjovsky Marc
- Page 276: 126 Dateor contracts put forth by t
- Page 280: 8Bioinformatics Tools for Modeling
- Page 284: Modeling Transcription Factor Targe
- Page 290: 134 DavuluriPWM-based models do not
- Page 294: 136 DavuluriTF-map alignments of or
- Page 298: 138 Davuluridiscussed which program
- Page 302: 140 DavuluriTable 2ER-a-Responsive
- Page 306: Table 3Sample Data Matrix Represent
- Page 310: Table 3 (Continued)Class MYCMAX MYC
- Page 314: 146 DavuluriFig. 3. (A) CART Tree:
- Page 318: 148 Davuluri11. Vlieghe, D., Sandel
- Page 322: 150 Davuluri44. Berezikov, E., Gury
- Page 326: 9Mining Biomedical Data Using MetaM
- Page 330: Mining Biomedical Data Using MMTx a
- Page 334: Mining Biomedical Data Using MMTx a
Table 1Web URLs of Promoter, TF Databases, and TFBs Prediction ProgramsProgram name Description Organism Web URL ReferencesTRANSFAC Database of TFs, their genomic, Eukaryotes http://www.gene-regulation.com 12binding sites and DNAbindingprofilesJASPAR Database of TF–DNA binding Eukaryotes http://mordor.cgb.ki.se/cgi- 11preferences, modeled bin/jaspar2005/jaspar_db.plas matricesMPromDb Database of mammalian Mammals http://bioinformatics.med.ohio- 15gene promoters state.edu/MPromDbOMGProm Database of orthologous Human, mouse, http://bioinformatics.med.ohio- 61mammalian gene promoters and rat state.edu/OMGPromDBTSS Database of transcription Human, mouse, http://dbtss.hgc.jp 16start sites zebrafish, malaria,and schyzonEPD Database of eukaryotic Eukaryotes http://www.epd.isb-sib.ch 67gene promotersUCSC Genome browser at UCSC Genome sequences http://genome.ucsc.edu 68and annotationsincluding human–mouse–ratconserved blocksMatch Program to search for TFBSs Eukaryotes http://www.gene-regulation.com/ 19using TRANSFAC PWMs pub/programs.html#matchTESS Program to predict TFBSs Eukaryotes http://www.cbil.upenn. –edu/cgi-bin/tess/tessMatInspector Program to search for TFBSs Eukaryotes http://www.genomatix.de/products/ 50using TRANSFAC PWMs MatInspector/MatInspector1.html132