Oncology Probes
Oncology Probes Oncology Probes
ON IGH (14q32) BreakChromosomal rearrangements involving the immunoglobulinheavy chain gene (IGH) at 14q32 are observed in 50% ofpatients with B-cell non-Hodgkin’s lymphoma (NHL) andmany other types of Lymphomas. More than 50 translocationpartners with IGH have been described. In particular t(8;14),is associated with Burkitt’s lymphoma, t(11;14) is associatedwith Mantle cell lymphoma, t(14;18) is observed in a highproportion of follicular lymphomas and t(3;14) is associatedwith Diffuse Large B-Cell Lymphoma.The IGH (14q32) break probe is optimized to detecttranslocations involving the IGH gene region at 14q32 in adual-color, split assay. Kreatech has developed this probe forthe specific use on cell material (KBI-10601), or for the useon tissue (KBI-10729).ON MALT (18q21) BreakLow grade malignant lymphomas arising from mucosaassociated lymphoid tissue (MALT) represent a distinctclinicopathological entity. The three major translocationsseen in MALT lymphomas are t(11;18)(q21;q21)/API2-MALT1,t(14;18)(q32;q21)/IGH-MALT1 and t(1;14)(p22;q32)/IGH-BCL10. A break or split probe for MALT (18q21) is best usedto analyze translocation of the MALT gene on formalin fixedparaffin embedded tissue for routine clinical diagnosis.Kreatech has developed this probe for the specific use on cellmaterial (KBI-10608), or for the use on tissue (KBI-10731).Cat.# KBI-10601 IGH (14q32) BreakCat.# KBI-10608 MALT (18q21) BreakD18S1055D14S683EJAG2345 KBIGHCIGHJIGHD14q32600 KB1818q21 MALTRH103885680 KB14IGHV400 KBD14S308Oncology Probes - Hematology ProbesIGH (14q32) Break probe hybridized to patient material showing apartial deletion of 14q32 (1RG1R).Literature:Taniwaki et al, 1994, Blood, 83: 2962-1969.Gozetti et al, 2002, Cancer Research, 62: 5523-5527.Ordering information Color Tests Cat#ON IGH (14q32) Break red/green 10 KBI-10601MALT (18q21) Break probe hybridized to patient material showing atranslocation at 18q21 (1RG1RG).Literature:Morgan et al, 1999, Cancer Res, 59; 6205-6213Dierlamm et al, 2000, Blood, 96; 2215-2218.Ordering information Color Tests Cat#ON MALT (18q21) Break red/green 10 KBI-1060842
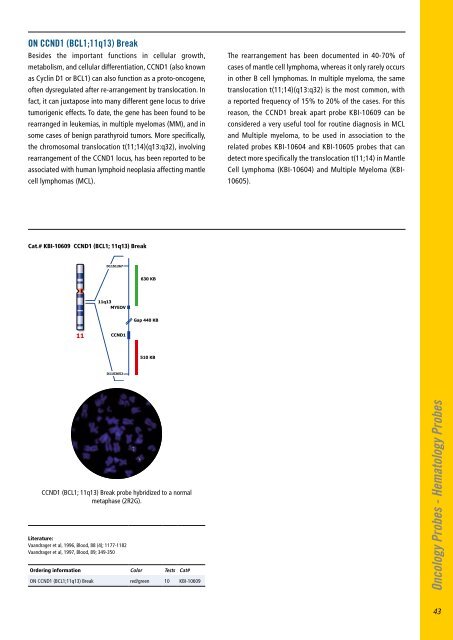
ON CCND1 (BCL1;11q13) BreakBesides the important functions in cellular growth,metabolism, and cellular differentiation, CCND1 (also knownas Cyclin D1 or BCL1) can also function as a proto-oncogene,often dysregulated after re-arrangement by translocation. Infact, it can juxtapose into many different gene locus to drivetumorigenic effects. To date, the gene has been found to berearranged in leukemias, in multiple myelomas (MM), and insome cases of benign parathyroid tumors. More specifically,the chromosomal translocation t(11;14)(q13:q32), involvingrearrangement of the CCND1 locus, has been reported to beassociated with human lymphoid neoplasia affecting mantlecell lymphomas (MCL).The rearrangement has been documented in 40-70% ofcases of mantle cell lymphoma, whereas it only rarely occursin other B cell lymphomas. In multiple myeloma, the sametranslocation t(11;14)(q13:q32) is the most common, witha reported frequency of 15% to 20% of the cases. For thisreason, the CCND1 break apart probe KBI-10609 can beconsidered a very useful tool for routine diagnosis in MCLand Multiple myeloma, to be used in association to therelated probes KBI-10604 and KBI-10605 probes that candetect more specifically the translocation t(11;14) in MantleCell Lymphoma (KBI-10604) and Multiple Myeloma (KBI-10605).Cat.# KBI-10609 CCND1 (BCL1; 11q13) BreakD11S1267630 KB11q13MYEOVGap 440 KB11CCND1510 KBD11S3652CCND1 (BCL1; 11q13) Break probe hybridized to a normalmetaphase (2R2G).Literature:Vaandrager et al, 1996, Blood, 88 (4); 1177-1182Vaandrager et al, 1997, Blood, 89; 349-350Ordering information Color Tests Cat#ON CCND1 (BCL1;11q13) Break red/green 10 KBI-10609Oncology Probes - Hematology Probes43
- Page 1: KREATECH DiagnosticsProduct Guide
- Page 4 and 5: Product Selection GuideI want to us
- Page 8 and 9: HUGO Gene SymbolsThe table below gi
- Page 10 and 11: Oncology probes - Hematology Probes
- Page 12 and 13: Chronic Myeloproliferative Disorder
- Page 14 and 15: ON BCR/ABL t(9;22), DC, S-Fusion, E
- Page 16 and 17: Other Myeloproliferative Diseases:O
- Page 18 and 19: Chronic Lymphocytic Leukemia (CLL)C
- Page 21: ON hTERC (3q26) / 3q11Amplification
- Page 25 and 26: ON hTERT (5p15) / 5q31The hTERT / 5
- Page 27 and 28: ON MDS 7q- (7q22; 7q35) / SE 7 TCTh
- Page 29 and 30: Acute Myeloid Leukemia (AML)At leas
- Page 31 and 32: ON RARA (17q21) BreakThis break apa
- Page 33 and 34: ON ETV6 (TEL) (12p13) BreakETV6 (TE
- Page 37 and 38: ON MM 1q21 / 8p21Amplifications of
- Page 39: ON MM 1q21 / SRD (1p36)Frequent los
- Page 44 and 45: ON BCL6 (3q27) BreakChromosomal tra
- Page 46 and 47: Oncology probes - Solid TumorsIn so
- Page 48 and 49: ON TOP2A (17q21) / Her2 (17q12) / S
- Page 50 and 51: Bladder CancerON p16 (9p21) / 9q21
- Page 52 and 53: NeuroblastomaAccording to the Inter
- Page 54 and 55: ON p53 (17p13) / MPO (17q22) “ISO
- Page 56 and 57: ON SYT (18q11) BreakThe characteris
- Page 58 and 59: ON MDM2 (12q15) / SE 12Fibrosarcoma
- Page 60 and 61: ON PTEN (10q23) / SE 10The gene ‘
- Page 62 and 63: ON CCND1 (11q13) / SE 11CCND1 (also
- Page 64 and 65: ON BCL6 (3q27) Break (tissue)Chromo
- Page 66 and 67: Preimplantation Genetic ScreeningFI
- Page 68 and 69: Prenatal probesPrenatal cytogenetic
- Page 70 and 71: Microdeletion probesMicrodeletion s
- Page 72 and 73: MD DiGeorge “TUPLE” (22q11) / 2
- Page 74 and 75: MD NSD1 (5q35) / hTERT (5p15)NSD1 m
- Page 76 and 77: MD Angelman UBE3A (15q11) / PML (15
- Page 78 and 79: MD Wolf-Hirschhorn WHSC1 (4p16)Wolf
- Page 80 and 81: MD STS (Xp22) / KAL (Xp22) / SE X T
- Page 82 and 83: Sub-Telomere probesTelomeres are sp
- Page 84 and 85: Satellite Enumeration probesThe pri
- Page 86 and 87: Whole Chromosome probesThe Poseidon
- Page 88 and 89: Arm Specific ProbesArm Specific Pro
- Page 90 and 91: Mouse FISH ProbesNew applications f
ON CCND1 (BCL1;11q13) BreakBesides the important functions in cellular growth,metabolism, and cellular differentiation, CCND1 (also knownas Cyclin D1 or BCL1) can also function as a proto-oncogene,often dysregulated after re-arrangement by translocation. Infact, it can juxtapose into many different gene locus to drivetumorigenic effects. To date, the gene has been found to berearranged in leukemias, in multiple myelomas (MM), and insome cases of benign parathyroid tumors. More specifically,the chromosomal translocation t(11;14)(q13:q32), involvingrearrangement of the CCND1 locus, has been reported to beassociated with human lymphoid neoplasia affecting mantlecell lymphomas (MCL).The rearrangement has been documented in 40-70% ofcases of mantle cell lymphoma, whereas it only rarely occursin other B cell lymphomas. In multiple myeloma, the sametranslocation t(11;14)(q13:q32) is the most common, witha reported frequency of 15% to 20% of the cases. For thisreason, the CCND1 break apart probe KBI-10609 can beconsidered a very useful tool for routine diagnosis in MCLand Multiple myeloma, to be used in association to therelated probes KBI-10604 and KBI-10605 probes that candetect more specifically the translocation t(11;14) in MantleCell Lymphoma (KBI-10604) and Multiple Myeloma (KBI-10605).Cat.# KBI-10609 CCND1 (BCL1; 11q13) BreakD11S1267630 KB11q13MYEOVGap 440 KB11CCND1510 KBD11S3652CCND1 (BCL1; 11q13) Break probe hybridized to a normalmetaphase (2R2G).Literature:Vaandrager et al, 1996, Blood, 88 (4); 1177-1182Vaandrager et al, 1997, Blood, 89; 349-350Ordering information Color Tests Cat#ON CCND1 (BCL1;11q13) Break red/green 10 KBI-10609<strong>Oncology</strong> <strong>Probes</strong> - Hematology <strong>Probes</strong>43