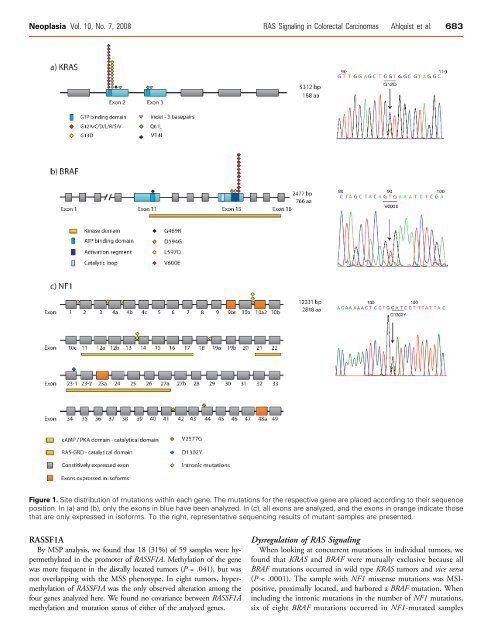
Neoplasia Vol. 10, No. 7, 2008 RAS Signal<strong>in</strong>g <strong>in</strong> Colorectal Carc<strong>in</strong>omas Ahlquist et al. 683Figure 1. Site distribution of mutations with<strong>in</strong> each gene. The mutations for the respective gene are placed accord<strong>in</strong>g to their sequenceposition. In (a) <strong>and</strong> (b), only the exons <strong>in</strong> blue have been analyzed. In (c), all exons are analyzed, <strong>and</strong> the exons <strong>in</strong> orange <strong>in</strong>dicate thosethat are only expressed <strong>in</strong> isoforms. To the right, representative sequenc<strong>in</strong>g results of mutant samples are presented.RASSF1ABy MSP analysis, we found that 18 (31%) of 59 samples were hypermethylated<strong>in</strong> the promoter of RASSF1A. Methylation of the genewas more frequent <strong>in</strong> the distally located tumors (P = .041), but was<strong>no</strong>t overlapp<strong>in</strong>g with the MSS phe<strong>no</strong>type. In eight tumors, hypermethylatio<strong>no</strong>f RASSF1A was the only observed alteration among thefour genes analyzed here. We found <strong>no</strong> covariance between RASSF1Amethylation <strong>and</strong> mutation status of either of the analyzed genes.Dysregulation of RAS Signal<strong>in</strong>gWhen look<strong>in</strong>g at concurrent mutations <strong>in</strong> <strong>in</strong>dividual tumors, wefound that KRAS <strong>and</strong> BRAF were mutually exclusive because allBRAF mutations occurred <strong>in</strong> wild type KRAS tumors <strong>and</strong> vice versa(P < .0001). The sample with NF1 missense mutations was MSIpositive,proximally located, <strong>and</strong> harbored a BRAF mutation. When<strong>in</strong>clud<strong>in</strong>g the <strong>in</strong>tronic mutations <strong>in</strong> the number of NF1 mutations,six of eight BRAF mutations occurred <strong>in</strong> NF1-mutated samples
684 RAS Signal<strong>in</strong>g <strong>in</strong> Colorectal Carc<strong>in</strong>omas Ahlquist et al. Neoplasia Vol. 10, No. 7, 2008(P = .03), overlapp<strong>in</strong>g with the MSI. Three of the four duplicationsfound <strong>in</strong> the NF1 locus occurred <strong>in</strong> tumors with wild typeBRAF <strong>and</strong> KRAS. The rema<strong>in</strong><strong>in</strong>g tumor had both a KRAS mutation<strong>and</strong> duplication.The occurrence of RASSF1A hypermethylation <strong>in</strong> the presence ofother mutations did <strong>no</strong>t show any trends toward coexistence or mutuallyexclusive nature.Taken together, we found that 74% (48/65) of the tumors had a<strong>no</strong>veractive RAS signal<strong>in</strong>g pathway due to change of at least one of thefour analyzed components (one alteration <strong>in</strong> 37/48, two <strong>alterations</strong> <strong>in</strong>10/48, <strong>and</strong> three <strong>alterations</strong> <strong>in</strong> 1/48). For the 24 samples submitted forcomplete analyses, the number of samples with at least one alterationwas 19 (79%): one alteration was seen <strong>in</strong> 14 samples, two <strong>alterations</strong>were seen <strong>in</strong> 4 samples, <strong>and</strong> three <strong>alterations</strong> were seen <strong>in</strong> 1 sample. Allsamples <strong>and</strong> <strong>alterations</strong> are summarized <strong>in</strong> Figure 2 <strong>and</strong> Table W2.DiscussionThis is the first report with an extensive analysis of the role of NF1mutations <strong>in</strong> colorectal tumorigenesis. Previous mutation studieshave only looked at a small number of samples, usually <strong>in</strong> a limitedpart of the gene, <strong>in</strong> the RAS–GAP doma<strong>in</strong>. The <strong>in</strong>itial mutationalreport on NF1 showed that 1 of 22 colorectal ade<strong>no</strong>carc<strong>in</strong>omas harboreda mutation <strong>in</strong> the RAS–GAP doma<strong>in</strong> us<strong>in</strong>g s<strong>in</strong>gle-str<strong>and</strong> conformationpolymorphism [9]. A<strong>no</strong>ther study of 10 colorectal celll<strong>in</strong>es <strong>and</strong> 4 sporadic tumors us<strong>in</strong>g prote<strong>in</strong> truncation test disclosedmutations <strong>in</strong> the NF1 cod<strong>in</strong>g region <strong>in</strong> four MSI cell l<strong>in</strong>es (40%)<strong>and</strong> one MSI tumor (25%). Two of the cell l<strong>in</strong>es had <strong>in</strong> fact twomutations each [22]. A recent study exam<strong>in</strong>ed five hereditary <strong>no</strong>npolyposisCRC patients for mutations <strong>in</strong> five exons <strong>and</strong> found a mutation<strong>in</strong> one (20%) of the patients who had a homozygous germl<strong>in</strong>emutation of MLH1 [23]. Moreover, loss of heterozygosity (LOH) atloci with<strong>in</strong> the NF1 gene have been shown <strong>in</strong> primary colorectal tumors(range, 14–57%) [24,25]. One of these studies also used realtimeexpression studies of NF1 <strong>in</strong> 55 of the carc<strong>in</strong>omas <strong>and</strong> found anFigure 2. Alterations across the sample series. The pie chart <strong>in</strong>dicatesthe four analyzed components <strong>and</strong> the percentage of tumorswhich showed <strong>alterations</strong> among these. Clockwise fromthe wild type pie, we see <strong>alterations</strong> <strong>in</strong> RASSF1A <strong>and</strong> BRAF; BRAF;BRAF <strong>and</strong> NF1; NF1; NF1 <strong>and</strong> KRAS; KRAS; KRAS <strong>and</strong> RASSF1A;RASSF1A; RASSF1A, NF1 <strong>and</strong> BRAF.<strong>in</strong>creased expression among tumors compared with <strong>no</strong>rmal colon tissue.In the COSMIC database [26], 79 carc<strong>in</strong>omas of the colorectumwere apparently <strong>in</strong>cluded among the NF1 data, yield<strong>in</strong>g a mutationfrequency of 11%. However, seven of n<strong>in</strong>e mutations reported werefrom one study <strong>in</strong>clud<strong>in</strong>g seven cell l<strong>in</strong>es, leav<strong>in</strong>g only two of the mutationsoccurr<strong>in</strong>g <strong>in</strong> sporadic primary tumors.In this study, we found the NF1 mutation profile to be <strong>in</strong> contrastto published germl<strong>in</strong>e mutation profile of NF1 patients 3 as well as tothe somatic mutation profiles of malignant peripheral nerve sheathtumor taken from patients with <strong>and</strong> without the NF1 disease [27,28](Bottillo I et al., unpublished observations). Furthermore, the medianage of the patients <strong>in</strong>cluded <strong>in</strong> the present CRC series is old, suggest<strong>in</strong>gthat potential NF1 carriers among them should have shown adebut of an NF1-associated cancer type. As <strong>no</strong> typical NF1-associatedtumors are recorded, based on written questionnaires <strong>and</strong> confirmatio<strong>no</strong>f cancer diag<strong>no</strong>ses from the Norwegian Cancer Registry [17],it further support that the reported mutations are somatic. The observed<strong>in</strong>tronic mutations prevail<strong>in</strong>g among the colorectal tumorscould be <strong>in</strong>volved <strong>in</strong> alternative splic<strong>in</strong>g but this rema<strong>in</strong>s to be elucidated.Four of the n<strong>in</strong>e <strong>in</strong>tronic mutations were <strong>in</strong>dels of one or twobases <strong>in</strong> microsatellites <strong>and</strong> reflect replication slippage (which often occurs<strong>in</strong> such repetitive stretches of bases) left unrepaired by the defectiveDNA mismatch repair system [29]. No such <strong>in</strong>dels were found <strong>in</strong>KRAS or BRAF.Multiple ligation-dependent probe amplification results showedthat 17% of the analyzed samples had ga<strong>in</strong>ed parts of or whole ofthe NF1 gene. This is <strong>no</strong>t <strong>in</strong> accordance with the expectations of atumor suppressor gene <strong>in</strong>volved <strong>in</strong> tumorigenesis. A duplication ofNF1 could lead to a stronger negative regulation of KRAS, with subsequentstronger control of proliferation <strong>and</strong> differentiation. The duplicationsmay arise as a consequence of the chromosomal <strong>in</strong>stabilitypresent <strong>in</strong> three of the four tumors, which yield a wide range of ga<strong>in</strong>s<strong>and</strong> losses of whole or parts of chromosomes. As reported by Ĉaĉevet al. [30], colorectal tumors show a significant <strong>in</strong>crease of NF1 mRNAexpression compared with correspond<strong>in</strong>g <strong>no</strong>rmal tissue. They alsoshowed that the expression of NF1 isoform I (lack<strong>in</strong>g exon 23a, located<strong>in</strong> the middle of the RAS-GRD doma<strong>in</strong>) was significantly higher <strong>in</strong>tumor compared with <strong>no</strong>rmal tissue [30]. As of this, the present f<strong>in</strong>d<strong>in</strong>gsare <strong>in</strong> agreement with those of the study by Ĉaĉev et al. [30].We also found a 40% mutation frequency of KRAS,whichiswith<strong>in</strong>the expected range [26]. A mutated KRAS (<strong>in</strong> codons 12, 13, <strong>and</strong> 61)h<strong>in</strong>ders the hydrolysis of GTP to GDP, <strong>and</strong> will keep KRAS <strong>in</strong> a constitutivelyactive state, lead<strong>in</strong>g to phosphorylation of downstream effectorssuch as BRAF [31]. BRAF mutations are k<strong>no</strong>wn to be stronglyassociated with MSI <strong>and</strong> CpG isl<strong>and</strong> methylator phe<strong>no</strong>type [32,33]<strong>and</strong> are found very often mutated <strong>in</strong> sessile-serrated ade<strong>no</strong>mas, lesionsoften considered as a precursor of MSI-H tumors [34–38]. We foundBRAF mutation <strong>in</strong> 22% of the samples, a higher frequency than <strong>in</strong>the mutation databases [26]. This reflects a bias due to the enrichmentof MSI tumors <strong>in</strong> the present series. In one study, 71% of theMSI tumors had a V600E mutation <strong>in</strong> BRAF, as opposed to 7% <strong>in</strong>the chromosomal-unstable tumors [39], a figure comparable with thepresent series, as 18 (62%) of 29 of MSI tumors had BRAF mutations.The most common BRAF mutation, V600E, just as the commonKRAS mutations, will lead to a constitutively active prote<strong>in</strong>, asthe activation loop of the prote<strong>in</strong> is changed [31].3 NF1 International Mutation Database (http://www.nfmutation.org).
- Page 1 and 2:
Novel genetic and epigenetic altera
- Page 3 and 4:
TABLE OF CONTENTSACKNOWLEDGEMENTS .
- Page 5 and 6:
ACKNOWLEDGEMENTSThe present work ha
- Page 7 and 8:
Prefacetechnology[3]. This new tech
- Page 10 and 11:
SummaryThe subgroup of carcinomas w
- Page 12 and 13:
Introduction“Epigenetic inheritan
- Page 14 and 15:
Introductionamino acid change it is
- Page 16 and 17:
Introductionmethylation during embr
- Page 18 and 19:
IntroductionDNA is most of the time
- Page 20 and 21:
IntroductionFigure 5. DNA methylati
- Page 22 and 23:
IntroductionFigure 6. Incidence rat
- Page 24 and 25:
IntroductionFigure 8. Tumor staging
- Page 26 and 27:
Introductioninasmuch as 80% of colo
- Page 28 and 29:
IntroductionInstabilities involved
- Page 30 and 31:
Introductionthere seems to be a fid
- Page 32 and 33:
Introductionsevere alterations are
- Page 34 and 35:
Introductionpopulation-wide screeni
- Page 36 and 37:
IntroductionFigure 12. Present and
- Page 38 and 39:
RESULTS IN BRIEFPaper Ia. “DNA hy
- Page 40 and 41:
Results in Briefinstability, and se
- Page 42 and 43:
Results in BriefUnivariate survival
- Page 44 and 45:
Discussionseveral factors, and full
- Page 46 and 47:
Discussionlow threshold, we increas
- Page 48 and 49:
DiscussionIt may seem like unnecess
- Page 50 and 51:
Discussionthan 96% DHPLC do not sta
- Page 52 and 53:
DiscussionFigure 13. Mutation detec
- Page 54 and 55:
DiscussionClinical impact of molecu
- Page 56 and 57:
Discussionmarkers with a very high
- Page 58 and 59:
Discussionchromosomes in metaphase[
- Page 60 and 61:
DiscussionThese examples underline
- Page 62 and 63:
Discussiongenes. One is based on mu
- Page 64 and 65:
CONCLUSIONSWe have identified novel
- Page 66 and 67:
Future PerspectivesMolecular risk a
- Page 68 and 69:
REFERENCES1. Breasted J (1930) The
- Page 70 and 71:
References29. Deng G, Chen A, Pong
- Page 72 and 73: References57. Al-Sukhni W, Aronson
- Page 74 and 75: References84. Kunkel TA (1993) Nucl
- Page 76 and 77: ReferencesLeggett B, Levine J, Kim
- Page 78 and 79: References133. Lind GE, Thorstensen
- Page 80 and 81: References156. Meling GI, Lothe RA,
- Page 82 and 83: ReferencesT, Song X, Day RH, Sledzi
- Page 84 and 85: References196. Honda S, Haruta M, S
- Page 86 and 87: ORIGINAL ARTICLESAPPENDIXAppendix I
- Page 89 and 90: GASTROENTEROLOGY 2007;132:1631-1639
- Page 91: Paper IbGuro E Lind, Terje Ahlquist
- Page 94 and 95: Journal of Translational Medicine 2
- Page 96 and 97: Journal of Translational Medicine 2
- Page 98 and 99: Journal of Translational Medicine 2
- Page 100 and 101: Journal of Translational Medicine 2
- Page 102 and 103: Journal of Translational Medicine 2
- Page 105: Paper IITerje Ahlquist, Guro E Lind
- Page 108 and 109: BackgroundMost cases of colorectal
- Page 110 and 111: ADAMTS1 CDKN2A CRABP1 HOXA9 MAL MGM
- Page 112 and 113: pseudogene, leading to a high rate
- Page 114 and 115: strands. Proc Natl Acad Sci U S A 1
- Page 116 and 117: concomitant absence of transcript a
- Page 119 and 120: Volume 10 Number 7 July 2008 pp. 68
- Page 121: 682 RAS Signaling in Colorectal Car
- Page 125 and 126: 686 RAS Signaling in Colorectal Car
- Page 127: Table W2. Detailed Somatic Events o
- Page 131 and 132: Identification of RCC2 as a prognos
- Page 133 and 134: INTRODUCTIONMicrosatellite instabil
- Page 135 and 136: unselected series of primary tumors
- Page 137 and 138: specificity, i.e. that they only am
- Page 139 and 140: On the assumption that DNA repair a
- Page 141 and 142: In order to ensure that gene mutati
- Page 143 and 144: Figure 2. Mutation frequency differ
- Page 145 and 146: and TAF1B (0.50), ACVR2A and ASTE1
- Page 147 and 148: Multivariate analysesA multivariate
- Page 149 and 150: When comparing our findings of muta
- Page 151 and 152: The test series included a low numb
- Page 153 and 154: entering M-phase remains to be seen
- Page 155 and 156: 12. Duval A, Reperant M, Hamelin R
- Page 157 and 158: 34. Martineau-Thuillier S, Andreass
- Page 159: AppendicesAppendix I:List of abbrev
- Page 163 and 164: Critical Reviews TM in Oncogenesis,
- Page 165 and 166: TARGET GENES OF MSI COLORECTAL CANC
- Page 167 and 168: TARGET GENES OF MSI COLORECTAL CANC
- Page 169 and 170: TARGET GENES OF MSI COLORECTAL CANC
- Page 171 and 172: TARGET GENES OF MSI COLORECTAL CANC
- Page 173 and 174:
TARGET GENES OF MSI COLORECTAL CANC
- Page 175 and 176:
TARGET GENES OF MSI COLORECTAL CANC
- Page 177 and 178:
TARGET GENES OF MSI COLORECTAL CANC
- Page 179 and 180:
TARGET GENES OF MSI COLORECTAL CANC
- Page 181 and 182:
TARGET GENES OF MSI COLORECTAL CANC
- Page 183 and 184:
TARGET GENES OF MSI COLORECTAL CANC
- Page 185 and 186:
TARGET GENES OF MSI COLORECTAL CANC
- Page 187 and 188:
TARGET GENES OF MSI COLORECTAL CANC
- Page 189 and 190:
TARGET GENES OF MSI COLORECTAL CANC
- Page 191:
TARGET GENES OF MSI COLORECTAL CANC