288 M. Hansson et al. / Developmental Biology 330 (2009) 286–304
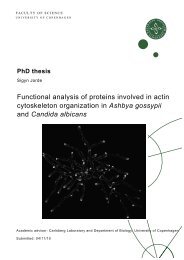
M. Hansson et al. / Developmental Biology 330 (2009) 286–304289specific secondary antibodies (Jackson ImmunoResearch Laboratories)and 4′,6-diamid<strong>in</strong>o-2-phenyl<strong>in</strong>dole (DAPI, MP Biomedicals).The Lhx1 sta<strong>in</strong><strong>in</strong>g used tyramide signal amplification (Perk<strong>in</strong>Elmer)accord<strong>in</strong>g to <strong>the</strong> manufacturer's recommendations. Negative controls,where <strong>the</strong> primary antibodies were omitted, were <strong>in</strong>cluded for allsta<strong>in</strong><strong>in</strong>gs. These controls showed no unspecific sta<strong>in</strong><strong>in</strong>g <strong>of</strong> <strong>the</strong>secondary antibodies (data not shown). β-galactosidase activity wasvisualized by add<strong>in</strong>g X-gal sta<strong>in</strong> solution for 4 h at 37 °C. The slideswere analyzed us<strong>in</strong>g an LSM 510 META laser scann<strong>in</strong>g microscope(Carl Zeiss) or a BX60 epifluorescence microscope equipped with aDP71 camera (both from Olympus).RT-PCR and qPCRCells were dissociated us<strong>in</strong>g 0.05% Tryps<strong>in</strong>-EDTA and collected bycentrifugation. Total RNA was isolated us<strong>in</strong>g <strong>the</strong> RNeasy kit withDNAse treatment (Qiagen) follow<strong>in</strong>g <strong>the</strong> manufacturer's protocol.cDNA was prepared from 5 or 100 ng RNA us<strong>in</strong>g MMLV ReverseTranscriptase (Invitrogen). PCR reactions were performed us<strong>in</strong>g 1 μlcDNA, 1 μl 20μM primer mix and 23 μl Reddy Mix PCR master mix(Abgene). The PCR was carried out with an <strong>in</strong>itial denaturation step at96 °C for 2 m<strong>in</strong>, followed by 32–38 cycles <strong>of</strong> 96 °C for 30 s, 55 °C for30 s and 72 °C for 1 m<strong>in</strong>. The PCR was f<strong>in</strong>ished with a f<strong>in</strong>al extensionstep at 72 °C for 5 m<strong>in</strong>. QPCR was performed us<strong>in</strong>g <strong>the</strong> standard SYBR ®Green program with dissociation curve <strong>of</strong> <strong>the</strong> Mx3005P (Stratagene).PCR reaction was run <strong>in</strong> duplicates us<strong>in</strong>g 5 μl Brilliant ® SYBR ® GreenQPCR Master Mix (Stratagene), 1 μl cDNA, 1 μl 10μM primer mix and3 μl DEPC-treated water. Quantified values for each gene <strong>of</strong> <strong>in</strong>terestwere normalized aga<strong>in</strong>st <strong>the</strong> <strong>in</strong>put determ<strong>in</strong>ed by <strong>the</strong> housekeep<strong>in</strong>ggenes G6pdh and Tbp. The results are expressed as <strong>the</strong> relativeexpression level compared with <strong>the</strong> vehicle control condition or <strong>the</strong>scrambled control siRNA <strong>in</strong> <strong>the</strong> vehicle condition. Primer sequencesare available on request.siRNA transfectionThe sequence effective for <strong>mouse</strong> β-caten<strong>in</strong> knock-down wasdesigned us<strong>in</strong>g s<strong>of</strong>tware available at <strong>the</strong> web site <strong>of</strong> Invitrogen(http://rnaidesigner.<strong>in</strong>vitrogen.com/rnaiexpress/). The β-caten<strong>in</strong>(accession number NM_0079614.2) target sequences <strong>of</strong> <strong>the</strong> STEALTHsiRNAs were 5′-GCCTTCATTATGGACTGCCTGTTGT-3′ (siRNA1) and 5′-GAGCAAGGCTTTTCCCAGTCCTTCA-3′ (siRNA2). The STEALTH negativecontrol siRNA (scrambled; Invitrogen) has been used as negativecontrol and is labeled “scrambled”.The <strong>cells</strong> were cultured on gelat<strong>in</strong>-treated 24-well plates aspreviously described with a start<strong>in</strong>g density <strong>of</strong> 4000 <strong>cells</strong>/cm 2 .After1 day <strong>of</strong> <strong>differentiation</strong> <strong>the</strong> <strong>cells</strong> were transfected with 100 nMSTEALTH siRNAs us<strong>in</strong>g Lip<strong>of</strong>ectam<strong>in</strong>e2000 accord<strong>in</strong>g to <strong>the</strong> manufacturer's<strong>in</strong>structions. The transfected <strong>cells</strong> were grown fur<strong>the</strong>r andanalyzed for β-caten<strong>in</strong> expression by western blot at days 3 and 5 <strong>of</strong><strong>differentiation</strong>. QPCR and flow cytometry analysis was performed atday 5.Western blotTotal cell lysates were obta<strong>in</strong>ed on days 3 and 5 <strong>of</strong> <strong>differentiation</strong>culture us<strong>in</strong>g RIPA lysis buffer. 20 μg <strong>of</strong> each prote<strong>in</strong> sample wereloaded and analyzed by western blot us<strong>in</strong>g a rabbit anti-β-caten<strong>in</strong>monoclonal antibody (Lab Vision) and a <strong>mouse</strong> anti-β-Act<strong>in</strong>monoclonal antibody (Sigma-Aldrich) as primary antibodies aswell as secondary HRP-conjugated antibodies (Santa CruzBiotechnology).Generation <strong>of</strong> stable Wnt-reporter <strong>ES</strong> cell l<strong>in</strong>es and chimera formationThe SuTOP-CFP construct was generated by cutt<strong>in</strong>g <strong>the</strong> luciferasegene from <strong>the</strong> Super8XTOPFLASH Wnt reporter (generously providedby Dr. R. Moon, University <strong>of</strong> Seattle, WA) with Fse and Nco1 andreplac<strong>in</strong>g it with Cerulean PCR product from <strong>the</strong> pmCerulean-C1vector (Rizzo et al., 2004). To generate stably transfected cell l<strong>in</strong>es, E14<strong>cells</strong> (Hooper et al., 1987) <strong>of</strong> low passage number were co-transfectedwith SuTOP-CFP and a ploxP-Neo vector, conferr<strong>in</strong>g resistance toNeomyc<strong>in</strong>. The relationship between plasmids was 10:1 (reporter:ploxP-Neo). Transfection was carried out us<strong>in</strong>g Lip<strong>of</strong>ectam<strong>in</strong>e2000(Invitrogen) and 24 h after transfection, selection was added (200 μg/ml G418, Invitrogen). Medium was changed daily for 9 days and0.5 μM BIO (GSK3β <strong>in</strong>hibitor, Calbiochem), which activates canonicalWnt signal<strong>in</strong>g, was added to <strong>the</strong> medium for <strong>the</strong> last 2 days <strong>of</strong>selection to identify Wnt-responsive colonies. Fluorescent colonieswere <strong>the</strong>n picked and expanded and one clone was selected forchimera formation via blastocyst <strong>in</strong>jection.E3.5 blastocysts were harvested by flush<strong>in</strong>g <strong>the</strong> uteri <strong>of</strong> mature,time mated NMRI mice (Taconic). Chimeric embryos were generatedby <strong>in</strong>jection <strong>of</strong> approximately 10 SuTOP-CFP <strong>ES</strong> <strong>cells</strong> <strong>in</strong>to <strong>the</strong>blastocoel us<strong>in</strong>g a paraff<strong>in</strong>-oil driven manual <strong>in</strong>jector (Cell TramVario, Eppendorf) and a Narishige micromanipulator. Follow<strong>in</strong>g 3 h <strong>of</strong>culture <strong>in</strong> M16 medium, embryos were transferred to <strong>the</strong> uterus <strong>of</strong>E2.5 pseudo-pregnant, 7 week old NMRI foster mo<strong>the</strong>rs. Care <strong>of</strong> <strong>the</strong>animals was done accord<strong>in</strong>g to <strong>in</strong>stitutional guidel<strong>in</strong>es. Embryos wereharvested at E10.5 and fixed <strong>in</strong> Lilly's fixative (4% phosphate bufferedformaldehyde) for 30 m<strong>in</strong> before be<strong>in</strong>g analyzed for native Ceruleanfluorescence. All animal experiments were performed <strong>in</strong> accordancewith <strong>in</strong>stitutional and national regulations.Chick embryo graft<strong>in</strong>gFertilized eggs from white leghorn chicken were purchased fromTriova and <strong>in</strong>cubated at 38 °C to Hamburger and Hamilton (HH) stages8–10 (Hamburger and Hamilton, 1951). The embryos were explantedas previously described (Chapman et al., 2001). E14 <strong>ES</strong> cell progenywas prepared for graft<strong>in</strong>g by label<strong>in</strong>g with fluorescent CMTMRCellTracker dye (Molecular Probes/Invitrogen). Clumps <strong>of</strong> <strong>cells</strong> werescraped <strong>of</strong>f, washed <strong>in</strong> PBS and <strong>in</strong>serted between endoderm andmesoderm <strong>of</strong> chicken embryos via a small <strong>in</strong>cision <strong>in</strong> <strong>the</strong> endoderm.Grafted embryos were <strong>in</strong>cubated for 48 h <strong>in</strong> a humidified <strong>in</strong>cubator at38 °C. The embryos were isolated, washed <strong>in</strong> PBS, fixed <strong>in</strong> 4% PFA atroom temperature for 2 h and stored <strong>in</strong> methanol at −20 °C until <strong>the</strong>time <strong>of</strong> analysis. Whole-mount immun<strong>of</strong>luorescent analyses <strong>of</strong>grafted chicken embryos were performed as previously described(Ahnfelt-Ronne et al., 2007).ResultsDose-dependent effects <strong>of</strong> activ<strong>in</strong> on <strong>the</strong> expression <strong>of</strong> PS genes aremodulated by BMP and Wnt signal<strong>in</strong>gPrevious studies have demonstrated <strong>in</strong>duction <strong>of</strong> Brachyury (T) andGoosecoid (Gsc) by activ<strong>in</strong> <strong>in</strong> m<strong>ES</strong> <strong>cells</strong> (Gadue et al., 2006; Kubo et al.,2004; Tada et al., 2005; Yasunaga et al., 2005). However, differences <strong>in</strong>media compositions and <strong>the</strong> culture methods used make a directcomparison <strong>of</strong> <strong>the</strong> response <strong>of</strong> <strong>the</strong>se two genes to vary<strong>in</strong>g doses <strong>of</strong>activ<strong>in</strong> difficult. Prior to <strong>the</strong> <strong>in</strong>duction <strong>of</strong> <strong>differentiation</strong> by addition <strong>of</strong>growth factors, we culture <strong>the</strong> undifferentiated <strong>ES</strong> <strong>cells</strong> under def<strong>in</strong>edconditions (Y<strong>in</strong>g et al., 2003a). As activ<strong>in</strong> is known to dosedependentlyregulate T and Gsc expression <strong>in</strong> Xenopus animal capFig. 1. Activ<strong>in</strong>, BMP and Wnt signal<strong>in</strong>g control <strong>the</strong> dynamic expression <strong>of</strong> primitive streak genes <strong>in</strong> differentiat<strong>in</strong>g <strong>ES</strong> <strong>cells</strong>. Flow cytometric analysis <strong>of</strong> T Gfp/+ (A), Gsc Gfp/+ (B), orMixl1 Gfp/+ (C) <strong>cells</strong> grown <strong>in</strong> adherent culture for up to 6 days <strong>in</strong> serum-free media conta<strong>in</strong><strong>in</strong>g 0, 3, 10, 30 or 100 ng/ml activ<strong>in</strong> <strong>in</strong> <strong>the</strong> presence or absence <strong>of</strong> 10 ng/ml BMP4 or100 ng/ml Wnt3a. The mean % GFP + <strong>cells</strong> ±standard deviation <strong>of</strong> three <strong>in</strong>dependent experiments is presented.
- Page 1: PhD thesisCand.scient. Janny Marie
- Page 5: ResuméSukkersyge er en sygdom der
- Page 9: Table of contents1
- Page 12 and 13: ICMinner cell massIdInhibitor of di
- Page 14 and 15: cell mass regenerates probably thro
- Page 16 and 17: Figure 1-1: Early embryo developmen
- Page 18 and 19: Figure 1-3: Regional expression of
- Page 20 and 21: The pluripotent stateThe pluripoten
- Page 25 and 26: There are four membrane-bound FGFRs
- Page 27: 2. AimsThe aim of this study was to
- Page 30 and 31: Developmental Biology 330 (2009) 28
- Page 34 and 35: 290 M. Hansson et al. / Development
- Page 36 and 37: 292 M. Hansson et al. / Development
- Page 38 and 39: 294 M. Hansson et al. / Development
- Page 40 and 41: 296 M. Hansson et al. / Development
- Page 42 and 43: 298 M. Hansson et al. / Development
- Page 44 and 45: 300 M. Hansson et al. / Development
- Page 46 and 47: 302 M. Hansson et al. / Development
- Page 48 and 49: 304 M. Hansson et al. / Development
- Page 50 and 51: Figure S2Figure S2: A subpopulation
- Page 52 and 53: Figure S4Figure S4: Expression of T
- Page 54 and 55: Figure S6Figure S6: qRT-PCR analyse
- Page 56 and 57: epithelium; Cdx2, expressed posteri
- Page 58 and 59: Figure 4-4: A high FGF4-concentrati
- Page 60 and 61: Figure 4-6: A 3-step protocol does
- Page 63 and 64: 5. Paper IIFGFR(IIIc)-activation in
- Page 65 and 66: AbstractProgress in embryonic stem
- Page 67 and 68: variants in their Ig-like domain II
- Page 69 and 70: Cell count and proliferation assayC
- Page 71 and 72: influence on the numbers of Sox17-G
- Page 73 and 74: undifferentiated cells, we found th
- Page 75 and 76: FGF4, 5, FGF8b and FGFR1, are expre
- Page 77 and 78: with EdU-stain (blue sample); and w
- Page 79 and 80: Olsen, S.K., J.Y. Li, C. Bromleigh,
- Page 81 and 82: FiguresFigure 1: Screen for FGFR-is
- Page 83 and 84:
Figure 3: Activation of FGFRb or FG
- Page 87 and 88:
Opposite, Figure 6: In the absence
- Page 89 and 90:
6. General discussionEndoderm diffe
- Page 91 and 92:
Overall, the multitude of FGF-signa
- Page 93 and 94:
transplantation is the spread of an
- Page 96:
AcknowledgementsThe work presented
- Page 99 and 100:
Chambers, I., D. Colby, M. Robertso
- Page 101 and 102:
Hawkins, V.J. Wroblewski, D.S. Li,
- Page 103 and 104:
Nishikawa, S.I., S. Nishikawa, M. H
- Page 105 and 106:
Tanimizu, N., H. Saito, K. Mostov,