articles - the School of Engineering and Applied Science - University ...
articles - the School of Engineering and Applied Science - University ...
articles - the School of Engineering and Applied Science - University ...
Create successful ePaper yourself
Turn your PDF publications into a flip-book with our unique Google optimized e-Paper software.
ARTICLES<br />
0.6<br />
0.3<br />
0.2<br />
n = 9<br />
0<br />
0.6<br />
0<br />
0.1<br />
0.3<br />
0.05<br />
18<br />
0<br />
0.6<br />
0<br />
Density<br />
0.3<br />
0.05<br />
28<br />
0<br />
0.6<br />
0<br />
0.05<br />
0.3<br />
0.025<br />
37<br />
0<br />
0.6<br />
0<br />
0.3<br />
0.025<br />
71<br />
0<br />
–45 –30 –15 0 15 30 45<br />
0<br />
–45 –30 –15 0 15 30 45<br />
Distance from <strong>the</strong> bilayer centre ( Å )<br />
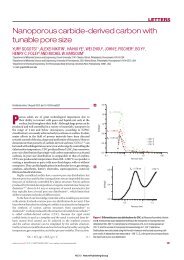
Figure 5 Density pr<strong>of</strong>iles revealing <strong>the</strong> chain entanglement <strong>and</strong> interdigitation.Density pr<strong>of</strong>iles <strong>of</strong> hydrophobic segments <strong>of</strong> <strong>the</strong> various polymersomes studied via CG-MD<br />
simulation.The number <strong>of</strong> hydrophobic segments (n ) in <strong>the</strong> corresponding copolymer is indicated in <strong>the</strong> figure.Left panel: Density pr<strong>of</strong>iles <strong>of</strong> hydrophobic blocks in each leaflet <strong>of</strong> <strong>the</strong><br />
corresponding bilayer are drawn in red <strong>and</strong> black,respectively.Right panel: Density pr<strong>of</strong>iles <strong>of</strong> <strong>the</strong> hydrophobic tail ends alone are shown.The left panel shows that <strong>the</strong> extent <strong>of</strong> overlap<br />
or interdigitation <strong>of</strong> hydrophobic segments increases with chain length <strong>and</strong>/or MW; <strong>the</strong> right panel demonstrates that <strong>the</strong> chain entanglement increases in <strong>the</strong> same direction.<br />
distinct regimes exist with an abrupt crossover at a MW phob ≈ 1.7 kDa.<br />
This provides an explanation for <strong>the</strong> experimental measurements <strong>of</strong><br />
copolymer diffusion in polymer vesicle membranes. The experiments<br />
likewise show a crossover from simple Rouse-like lateral mobility to a<br />
more complicated activated reptation for hydrophobic molecular<br />
weights > 2 kDa (refs 47,48).<br />
To generate fur<strong>the</strong>r insight into <strong>the</strong> entanglement mechanism,<br />
configurations <strong>of</strong> individual copolymer chains in <strong>the</strong> simulation were<br />
examined.A typical snapshot <strong>of</strong> two entangled copolymers is shown in<br />
Fig. 6b. Such entanglements are not observed with <strong>the</strong> smaller-MW<br />
block copolymers where <strong>the</strong> chains appear short, stiff <strong>and</strong> straight.<br />
The difference suggests that <strong>the</strong> observed trends in diffusional dynamics<br />
Table 1 Description <strong>of</strong> various copolymers studied in this work.The number <strong>of</strong> water sites (N water ) needed to hydrate a specific number <strong>of</strong> copolymers (N poly ) increases with<br />
increasing chain length.The hydrophilic fraction in each chain is represented by f phil .The molecular weight <strong>of</strong> each chain is denoted by MW while <strong>the</strong> respective hydrophobic<br />
mass is represented by MW phob. The hydrophobic core thickness (d ) <strong>and</strong> area per polymer chain (A ) are listed only for pre-assembled bilayer structures.For <strong>the</strong> worm <strong>and</strong><br />
spherical structures d denotes <strong>the</strong> diameter.<br />
Copolymer N poly /N water f phil MW/MW phob A Morphology d<br />
(%) (kDa) (Å 2 ) (nm)<br />
EO 10 EE 9 108/2,160 45.6% 1.03/0.50 67.07 Bilayer 2.63<br />
EO 19 EE 18 128/4,000 44.8% 1.93/1.00 76.10 Bilayer 4.38<br />
EO 29 EE 28 128/4,265 44.5% 2.93/1.57 94.34 Bilayer 5.99<br />
EO 40 EE 37 128/7,776 45.6% 3.92/2.07 104.2 Bilayer 7.68<br />
EO 57 EE 60 96/8,520 42.6% 5.96/3.36 134.3 Bilayer 8.66<br />
EO 73 EE 71 96/10,368 44.5% 7.28/3.98 146.4 Bilayer 9.79<br />
EO 21 EE 37 100/7,000 30.9% 3.08/2.07 - Bilayer 8.55<br />
EO 50 EE 37 64/7,680 51.1% 4.36/2.07 - Worm 6.01<br />
EO 92 EE 37 48/24,000 65.6% 6.21/2.07 - Spherical 4.83<br />
nature materials | ADVANCE ONLINE PUBLICATION | www.nature.com/naturematerials 5<br />
© 2004 Nature Publishing Group