pdf - A test of psbK-psbI and atpF-atpH as potential plant DNA - ArBOL
pdf - A test of psbK-psbI and atpF-atpH as potential plant DNA - ArBOL
pdf - A test of psbK-psbI and atpF-atpH as potential plant DNA - ArBOL
You also want an ePaper? Increase the reach of your titles
YUMPU automatically turns print PDFs into web optimized ePapers that Google loves.
Nature Precedings : hdl:10101/npre.2008.1896.1 : Posted 16 May 2008<br />
In a multi loci approach for <strong>DNA</strong> barcoding purposes, the highest mean <strong>of</strong> inter-specific<br />
variability w<strong>as</strong> achieved by matK combined with trnH-psbA <strong>and</strong> <strong>atpF</strong>-<strong>atpH</strong> where<strong>as</strong> the<br />
highest mean <strong>of</strong> intra-specific distances were given by combining matK with trnH-psbA<br />
(Table 3). Wilcoxon statistical rank <strong>test</strong>s showed the combination matK + trnH-psbA<br />
having the highest inter-specific pair-distances (Table 4). They revealed also that all the<br />
combinations including trnH-psbA had a higher intra-specific variability than<br />
combinations without it (Table 5).<br />
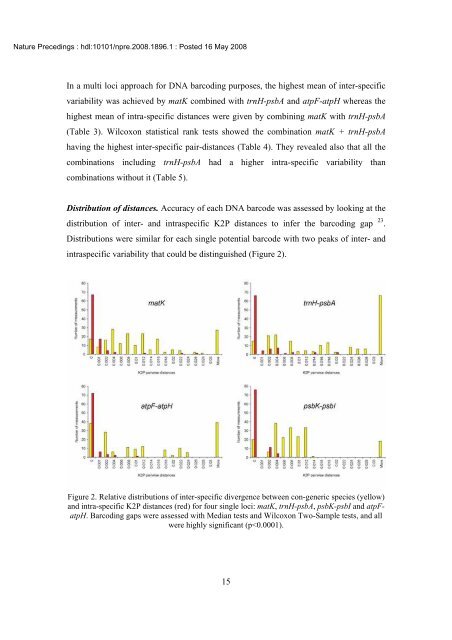
Distribution <strong>of</strong> distances. Accuracy <strong>of</strong> each <strong>DNA</strong> barcode w<strong>as</strong> <strong>as</strong>sessed by looking at the<br />
distribution <strong>of</strong> inter- <strong>and</strong> intr<strong>as</strong>pecific K2P distances to infer the barcoding gap 23 .<br />
Distributions were similar for each single <strong>potential</strong> barcode with two peaks <strong>of</strong> inter- <strong>and</strong><br />
intr<strong>as</strong>pecific variability that could be distinguished (Figure 2).<br />
Figure 2. Relative distributions <strong>of</strong> inter-specific divergence between con-generic species (yellow)<br />
<strong>and</strong> intra-specific K2P distances (red) for four single loci: matK, trnH-psbA, <strong>psbK</strong>-<strong>psbI</strong> <strong>and</strong> <strong>atpF</strong><strong>atpH</strong>.<br />
Barcoding gaps were <strong>as</strong>sessed with Median <strong>test</strong>s <strong>and</strong> Wilcoxon Two-Sample <strong>test</strong>s, <strong>and</strong> all<br />
were highly significant (p