Voie d'immunisation et séquence d'administration de l ... - TEL
Voie d'immunisation et séquence d'administration de l ... - TEL Voie d'immunisation et séquence d'administration de l ... - TEL
tel-00827710, version 1 - 29 May 2013 demonstrated that they are highly specific for their cognate ligand and, upon engagement, different TLRs induce a variety of signaling pathways (Figure 5). While TLR3 and TLR4 signal through the adaptor molecule TRIF and trigger the production of type I IFN and inflammatory cytokines, TLR1, 2, 4, 5 and 6 induce only inflammatory cytokines via signaling through Myd88-dependent pathways (Kawai and Akira, 2010). TLR 7 and 9 also signal via Myd88. Figure 5. Signaling pathways downstream of TLRs. (A) Myd88-dependent pathway downstream of TLR1/6, 2 and 4 and TRIF-dependent pathway activated by TLR3 and 4. (B) Myd88-dependent pathway downstream of TLR7 and 9. Adapted from Takeuchi et al., 2010. (ii) RIG-I-Like Receptors The RIG-I-like receptors are critical for the detection of infecting viruses. Three cytoplasmic receptors form this family. RIG-I, Mda5 and LPG2 are specific for viral RNA species (Table 2). RIG-I, in particular, detects short double-stranded RNAs only found during the replication of RNA viruses, while Mda5 recognizes longer nucleic acid chains. (iii) Nod-Like Receptors The NOD-like receptor family consists of more than 20 unique members that are all cytosolic (Table 2). Among them, NOD1 and NOD2 recognize the degradation products of bacterial 32
tel-00827710, version 1 - 29 May 2013 wall components. NLRP3 (NALP3) responds to various stimuli as part of the recently characterized inflammasome complex, which results in the cleavage of pro-IL-1β and pro-IL- 18 via the activation of Caspase 1. Table 2. Diversity of RLRs, NLRs, and CLRs. ssRNA, single stranded RNA; dsRNA, double stranded RNA; iE-DAP, dipeptide present in bacterial peptidoglycan; MDP, muramyl dipeptide. (iv) CLRs CLRs are transmembrane receptors that recognize carbohydrates from microorganisms (Table 2). As an example, Dectin-1 and Dectin-2 detect β-glucans from fungi. Clec9A is the prototypic member of this family. It is expressed on CD8α + DCs and is responsible for recognizing necrotic cells (Takeuchi and Akira, 2010). Page 33 of 256
- Page 1 and 2: tel-00827710, version 1 - 29 May 20
- Page 3 and 4: tel-00827710, version 1 - 29 May 20
- Page 5 and 6: tel-00827710, version 1 - 29 May 20
- Page 7 and 8: tel-00827710, version 1 - 29 May 20
- Page 9 and 10: tel-00827710, version 1 - 29 May 20
- Page 11 and 12: tel-00827710, version 1 - 29 May 20
- Page 13 and 14: tel-00827710, version 1 - 29 May 20
- Page 15 and 16: tel-00827710, version 1 - 29 May 20
- Page 17 and 18: tel-00827710, version 1 - 29 May 20
- Page 19 and 20: tel-00827710, version 1 - 29 May 20
- Page 21 and 22: tel-00827710, version 1 - 29 May 20
- Page 23 and 24: tel-00827710, version 1 - 29 May 20
- Page 25 and 26: tel-00827710, version 1 - 29 May 20
- Page 27 and 28: tel-00827710, version 1 - 29 May 20
- Page 29 and 30: tel-00827710, version 1 - 29 May 20
- Page 31: tel-00827710, version 1 - 29 May 20
- Page 35 and 36: tel-00827710, version 1 - 29 May 20
- Page 37 and 38: tel-00827710, version 1 - 29 May 20
- Page 39 and 40: tel-00827710, version 1 - 29 May 20
- Page 41 and 42: tel-00827710, version 1 - 29 May 20
- Page 43 and 44: tel-00827710, version 1 - 29 May 20
- Page 45 and 46: tel-00827710, version 1 - 29 May 20
- Page 47 and 48: tel-00827710, version 1 - 29 May 20
- Page 49 and 50: tel-00827710, version 1 - 29 May 20
- Page 51 and 52: tel-00827710, version 1 - 29 May 20
- Page 53 and 54: tel-00827710, version 1 - 29 May 20
- Page 55 and 56: tel-00827710, version 1 - 29 May 20
- Page 57 and 58: tel-00827710, version 1 - 29 May 20
- Page 59 and 60: tel-00827710, version 1 - 29 May 20
- Page 61 and 62: tel-00827710, version 1 - 29 May 20
- Page 63 and 64: tel-00827710, version 1 - 29 May 20
- Page 65 and 66: tel-00827710, version 1 - 29 May 20
- Page 67 and 68: tel-00827710, version 1 - 29 May 20
- Page 69 and 70: tel-00827710, version 1 - 29 May 20
- Page 71 and 72: tel-00827710, version 1 - 29 May 20
- Page 73 and 74: tel-00827710, version 1 - 29 May 20
- Page 75 and 76: tel-00827710, version 1 - 29 May 20
- Page 77 and 78: tel-00827710, version 1 - 29 May 20
- Page 79 and 80: tel-00827710, version 1 - 29 May 20
- Page 81 and 82: tel-00827710, version 1 - 29 May 20
tel-00827710, version 1 - 29 May 2013<br />
<strong>de</strong>monstrated that they are highly specific for their cognate ligand and, upon engagement,<br />
different TLRs induce a vari<strong>et</strong>y of signaling pathways (Figure 5). While TLR3 and TLR4<br />
signal through the adaptor molecule TRIF and trigger the production of type I IFN and<br />
inflammatory cytokines, TLR1, 2, 4, 5 and 6 induce only inflammatory cytokines via<br />
signaling through Myd88-<strong>de</strong>pen<strong>de</strong>nt pathways (Kawai and Akira, 2010). TLR 7 and 9 also<br />
signal via Myd88.<br />
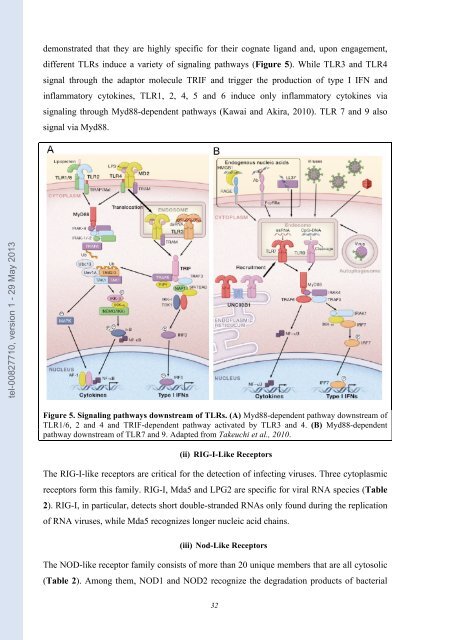
Figure 5. Signaling pathways downstream of TLRs. (A) Myd88-<strong>de</strong>pen<strong>de</strong>nt pathway downstream of<br />
TLR1/6, 2 and 4 and TRIF-<strong>de</strong>pen<strong>de</strong>nt pathway activated by TLR3 and 4. (B) Myd88-<strong>de</strong>pen<strong>de</strong>nt<br />
pathway downstream of TLR7 and 9. Adapted from Takeuchi <strong>et</strong> al., 2010.<br />
(ii) RIG-I-Like Receptors<br />
The RIG-I-like receptors are critical for the d<strong>et</strong>ection of infecting viruses. Three cytoplasmic<br />
receptors form this family. RIG-I, Mda5 and LPG2 are specific for viral RNA species (Table<br />
2). RIG-I, in particular, d<strong>et</strong>ects short double-stran<strong>de</strong>d RNAs only found during the replication<br />
of RNA viruses, while Mda5 recognizes longer nucleic acid chains.<br />
(iii) Nod-Like Receptors<br />
The NOD-like receptor family consists of more than 20 unique members that are all cytosolic<br />
(Table 2). Among them, NOD1 and NOD2 recognize the <strong>de</strong>gradation products of bacterial<br />
32